Identifying common transcriptome signatures of cancer by interpreting deep learning models, Genome Biology
Por um escritor misterioso
Last updated 13 junho 2024
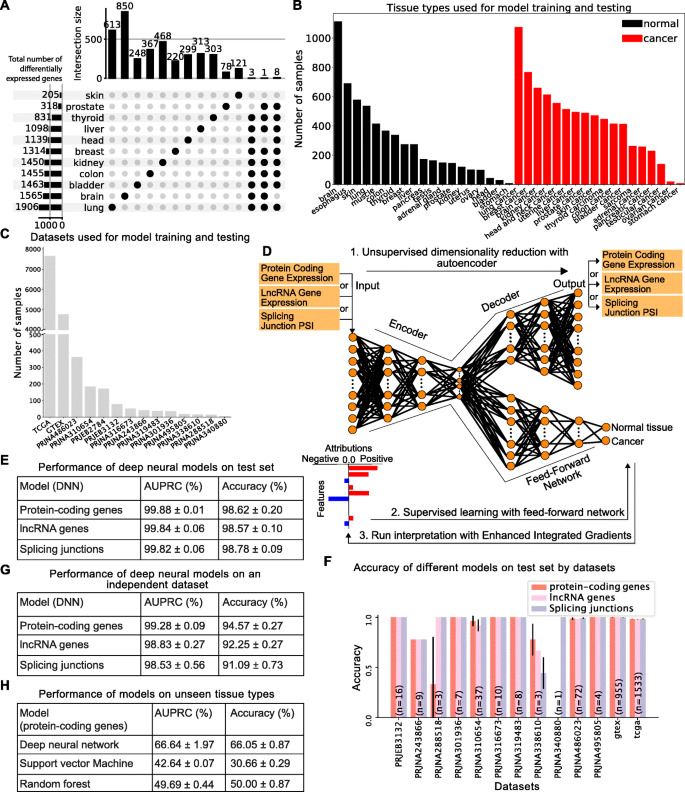
Background Cancer is a set of diseases characterized by unchecked cell proliferation and invasion of surrounding tissues. The many genes that have been genetically associated with cancer or shown to directly contribute to oncogenesis vary widely between tumor types, but common gene signatures that relate to core cancer pathways have also been identified. It is not clear, however, whether there exist additional sets of genes or transcriptomic features that are less well known in cancer biology but that are also commonly deregulated across several cancer types. Results Here, we agnostically identify transcriptomic features that are commonly shared between cancer types using 13,461 RNA-seq samples from 19 normal tissue types and 18 solid tumor types to train three feed-forward neural networks, based either on protein-coding gene expression, lncRNA expression, or splice junction use, to distinguish between normal and tumor samples. All three models recognize transcriptome signatures that are consistent across tumors. Analysis of attribution values extracted from our models reveals that genes that are commonly altered in cancer by expression or splicing variations are under strong evolutionary and selective constraints. Importantly, we find that genes composing our cancer transcriptome signatures are not frequently affected by mutations or genomic alterations and that their functions differ widely from the genes genetically associated with cancer. Conclusions Our results highlighted that deregulation of RNA-processing genes and aberrant splicing are pervasive features on which core cancer pathways might converge across a large array of solid tumor types.
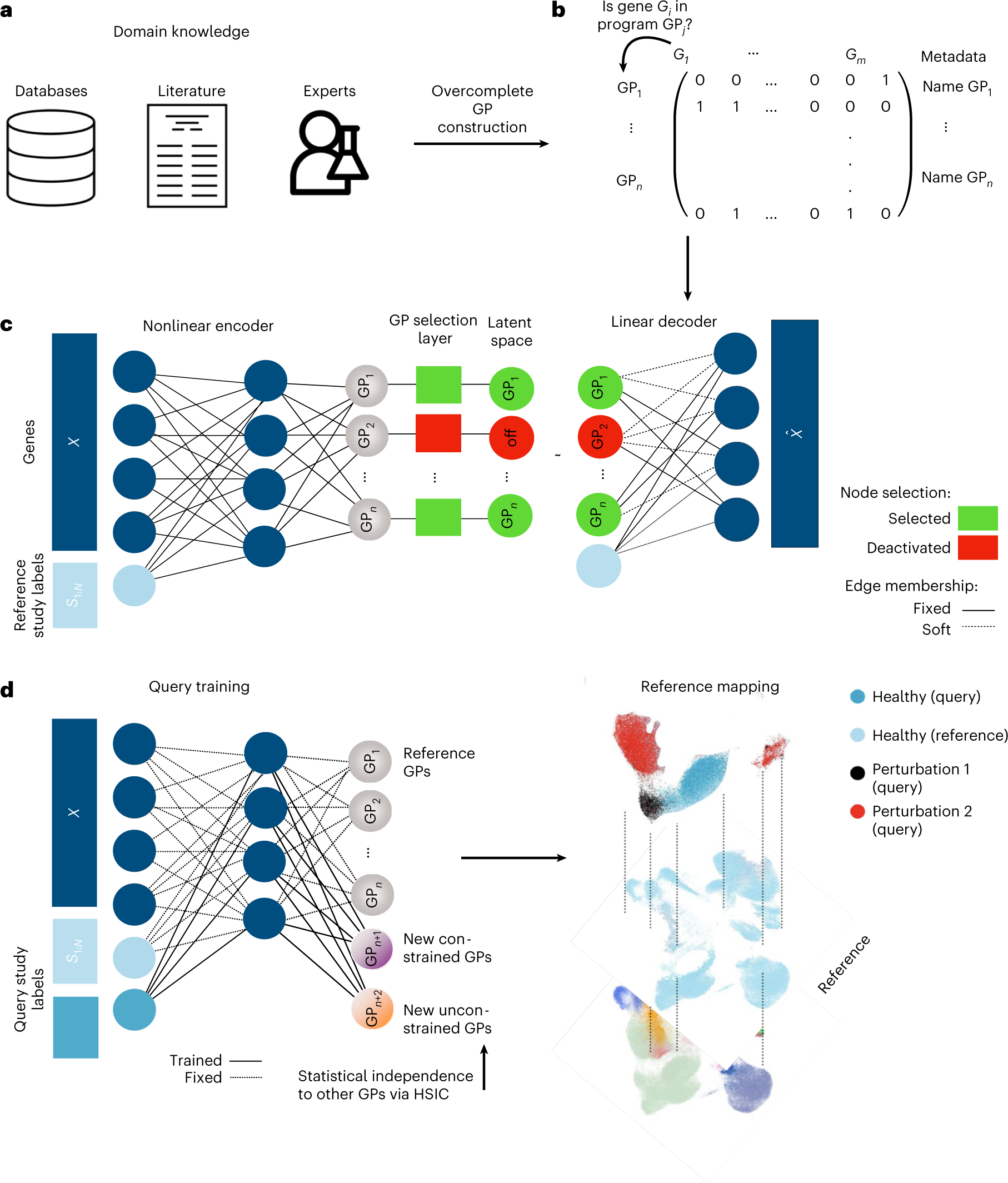
Biologically informed deep learning to query gene programs in
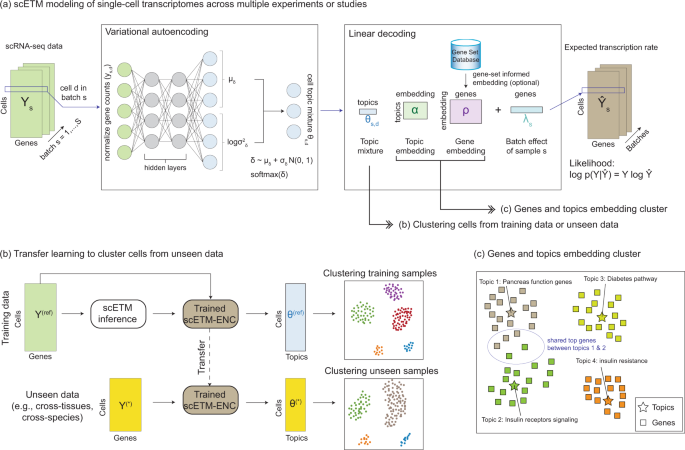
Learning interpretable cellular and gene signature embeddings from
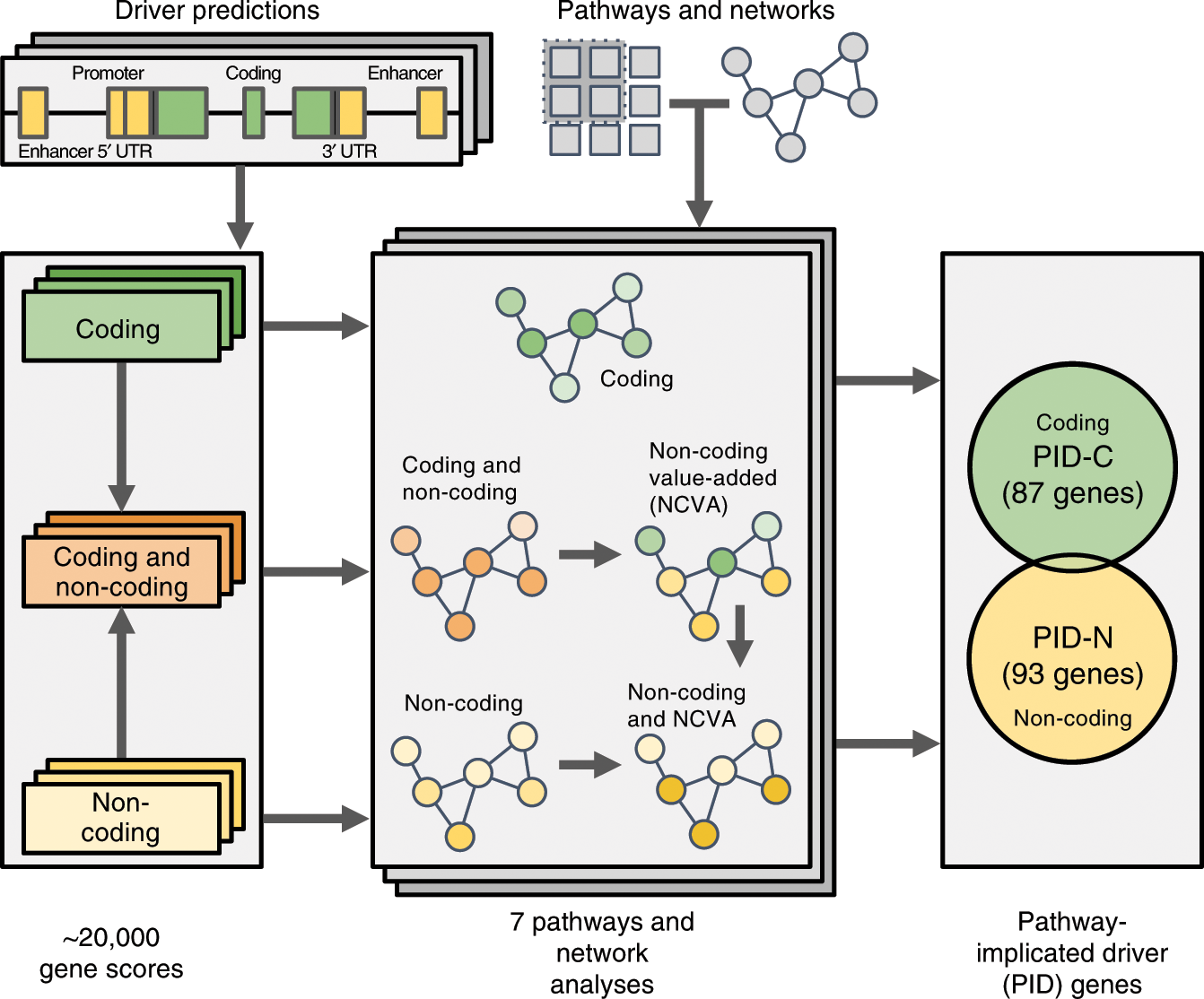
Pathway and network analysis of more than 2500 whole cancer
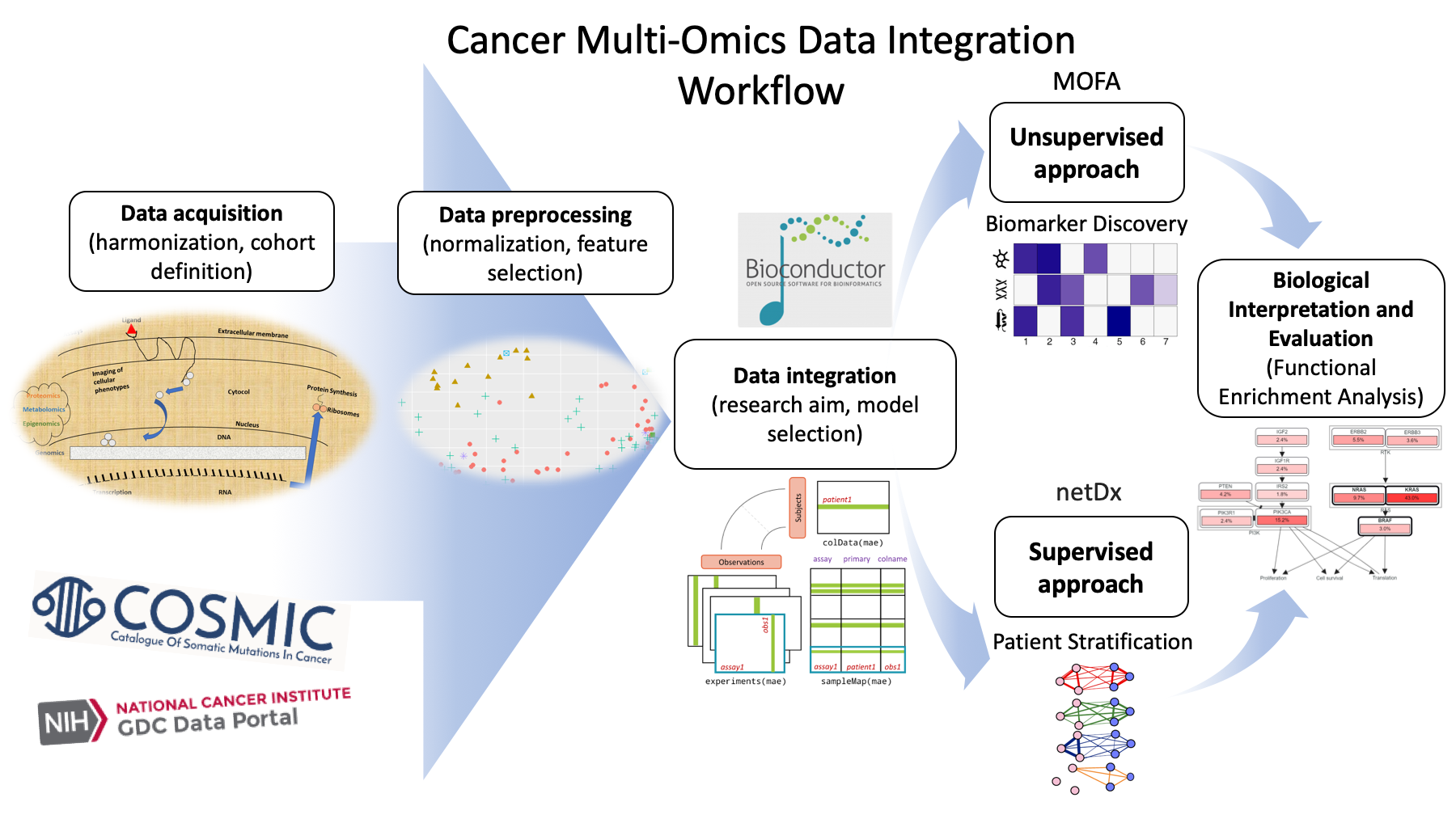
IJMS, Free Full-Text
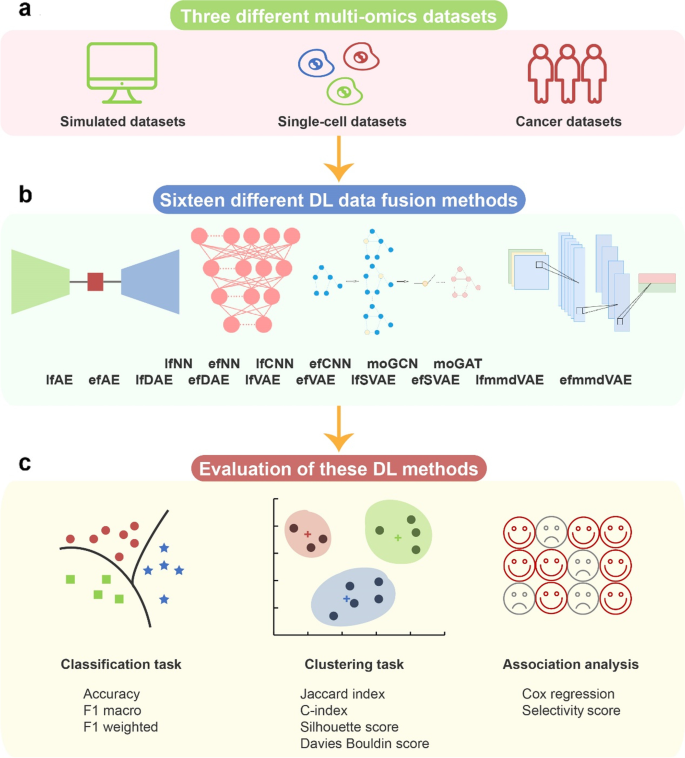
A benchmark study of deep learning-based multi-omics data fusion
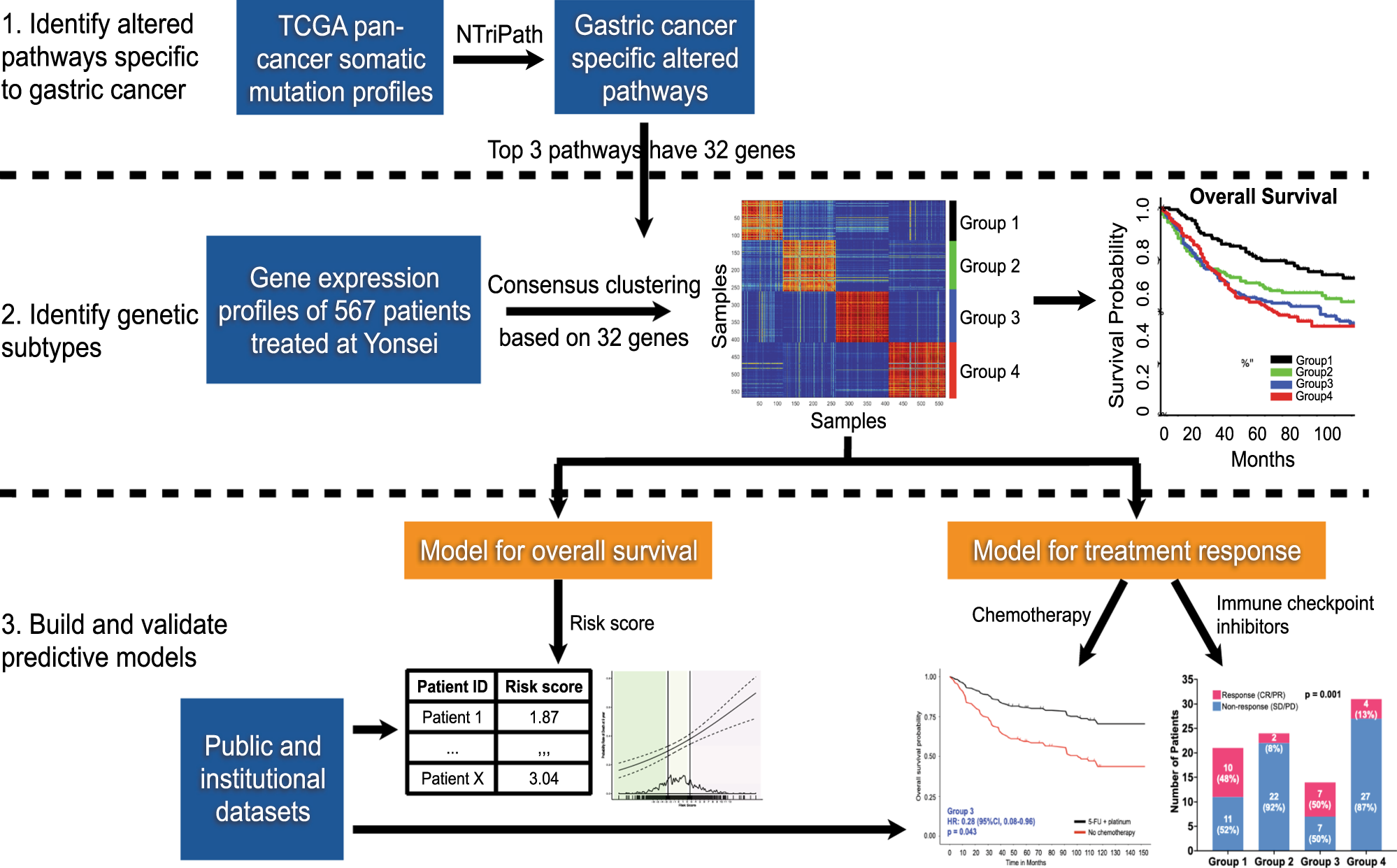
Development and validation of a prognostic and predictive 32-gene
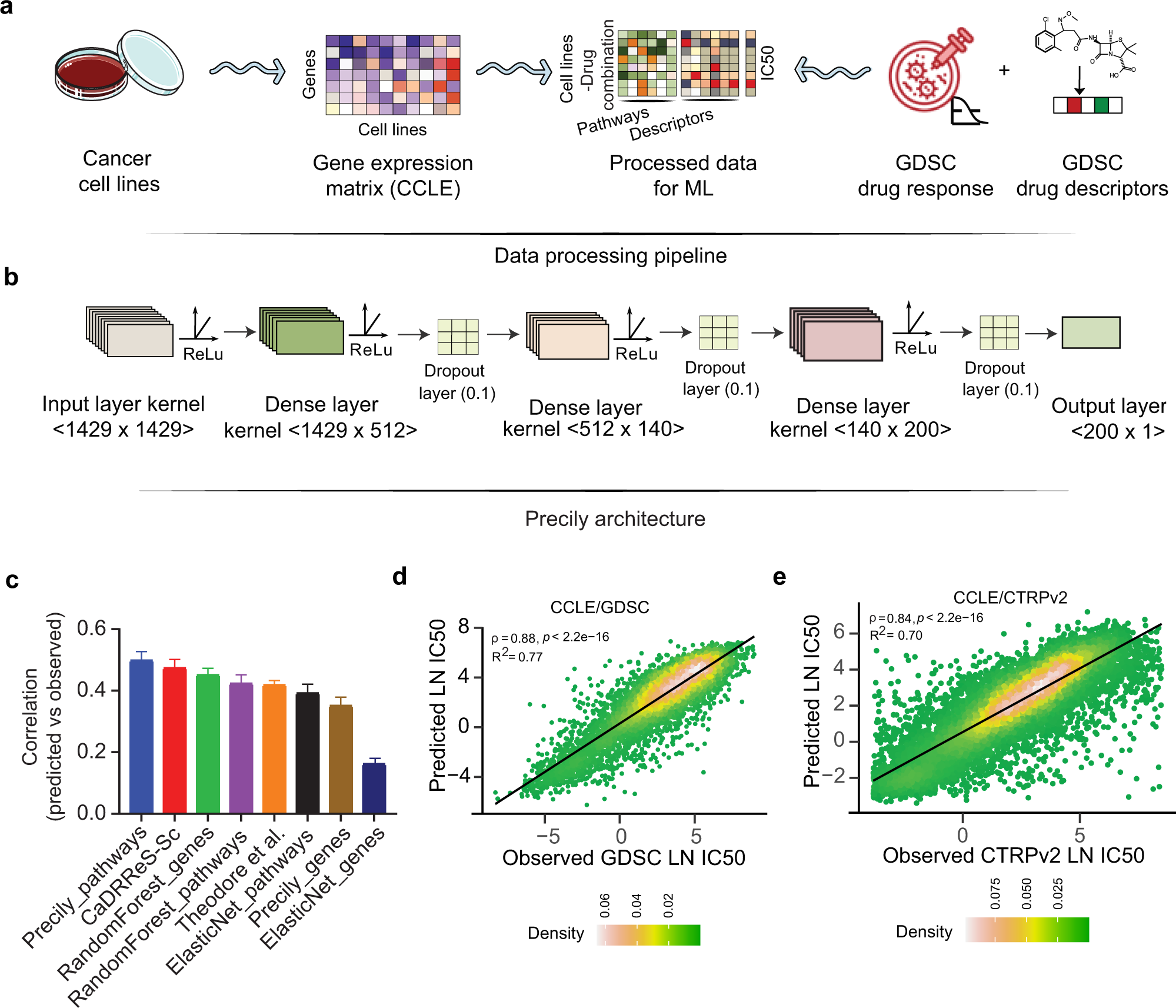
Gene expression based inference of cancer drug sensitivity
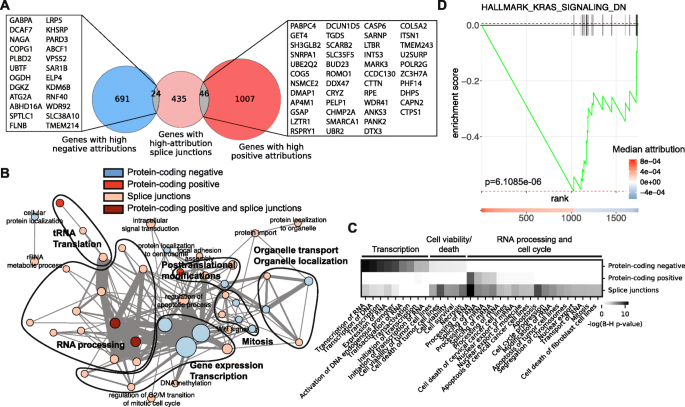
Identifying common transcriptome signatures of cancer by
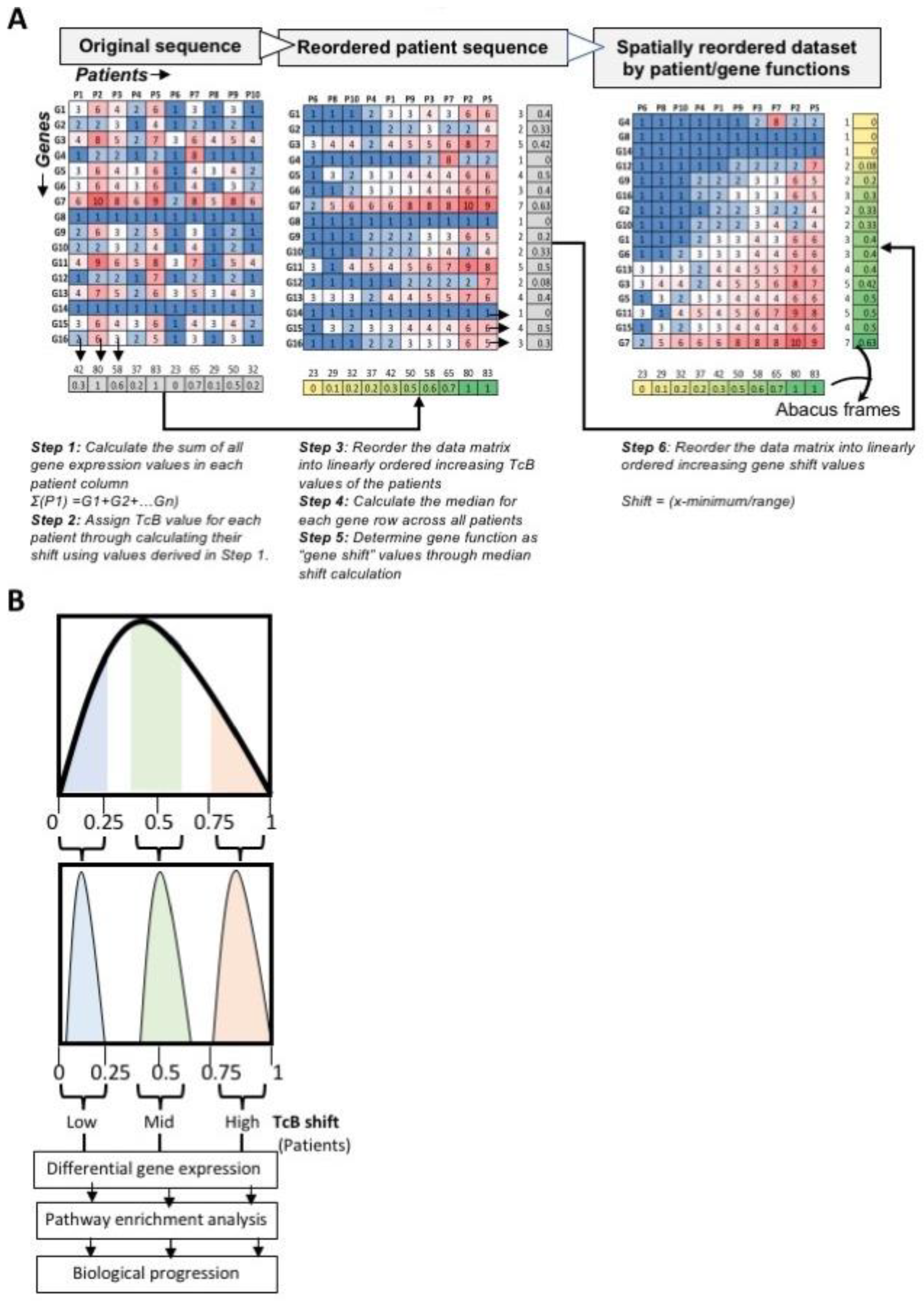
Biomedicines, Free Full-Text

Identifying common transcriptome signatures of cancer by
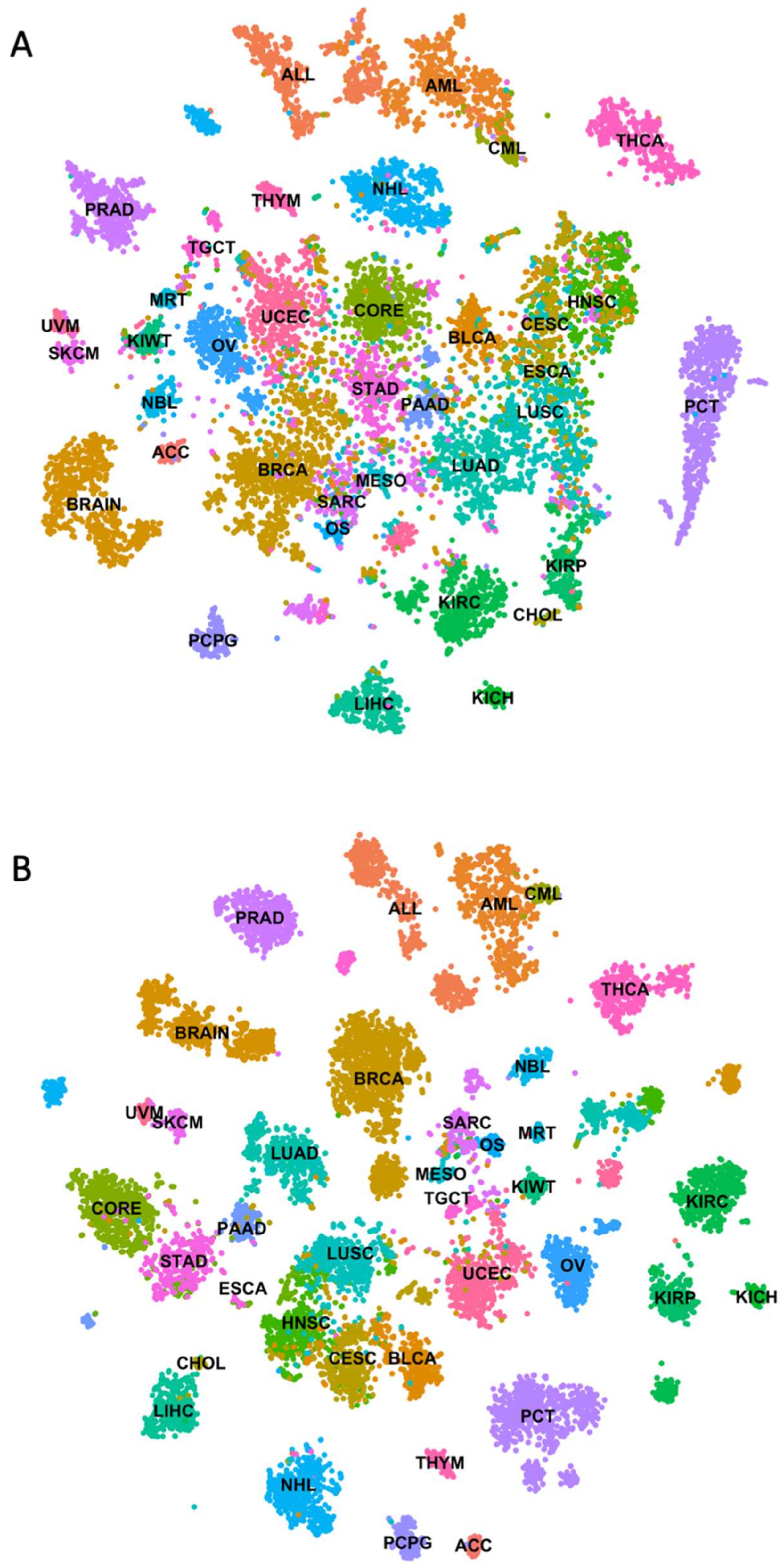
Cancers, Free Full-Text
Recomendado para você
-
 Brain Test Level 367 answer/solution. #shorts #braintest13 junho 2024
Brain Test Level 367 answer/solution. #shorts #braintest13 junho 2024 -
 Brain Test Level 372 He wants big muscles in 202313 junho 2024
Brain Test Level 372 He wants big muscles in 202313 junho 2024 -
 Results of whole brain analyses in the test scan. a The left parietal13 junho 2024
Results of whole brain analyses in the test scan. a The left parietal13 junho 2024 -
![PDF] Low-Dimensional Structure in the Space of Language Representations is Reflected in Brain Responses](https://d3i71xaburhd42.cloudfront.net/89711a7e91ad81874a5a0b1c3359bb67b27c0578/15-Table1-1.png) PDF] Low-Dimensional Structure in the Space of Language Representations is Reflected in Brain Responses13 junho 2024
PDF] Low-Dimensional Structure in the Space of Language Representations is Reflected in Brain Responses13 junho 2024 -
 Ryuta Kawashima: The devil who cracked the dementia code, The Independent13 junho 2024
Ryuta Kawashima: The devil who cracked the dementia code, The Independent13 junho 2024 -
 HOME13 junho 2024
HOME13 junho 2024 -
 Can Spinal Cord Injuries Affect the Brain? - Total Community Care13 junho 2024
Can Spinal Cord Injuries Affect the Brain? - Total Community Care13 junho 2024 -
 Gone West: Boeing Test Pilot James Gannett13 junho 2024
Gone West: Boeing Test Pilot James Gannett13 junho 2024 -
 Gut Microbiome–Brain Alliance: A Landscape View into Mental and13 junho 2024
Gut Microbiome–Brain Alliance: A Landscape View into Mental and13 junho 2024 -
 Brain Test Level 361, 362, 363, 364, 365, 366, 367, 368, 369, 37013 junho 2024
Brain Test Level 361, 362, 363, 364, 365, 366, 367, 368, 369, 37013 junho 2024
você pode gostar
-
 Fantasy Bishoujo Juniku Ojisan to – 03 – Random Curiosity13 junho 2024
Fantasy Bishoujo Juniku Ojisan to – 03 – Random Curiosity13 junho 2024 -
 🔥STUMBLE GUYS AO VIVO 🔥 - VEM JOGAR COMIGO !!🔥LIVE ON🔥13 junho 2024
🔥STUMBLE GUYS AO VIVO 🔥 - VEM JOGAR COMIGO !!🔥LIVE ON🔥13 junho 2024 -
Tic Tac Toe 5x5 - Apps on Google Play13 junho 2024
-
 Rain World's Downpour DLC to include 4-player local co-op13 junho 2024
Rain World's Downpour DLC to include 4-player local co-op13 junho 2024 -
 r63 roblox pause|TikTok Search13 junho 2024
r63 roblox pause|TikTok Search13 junho 2024 -
 Projeto Xeque Mate: conheça os benefícios do xadrez - Instituto13 junho 2024
Projeto Xeque Mate: conheça os benefícios do xadrez - Instituto13 junho 2024 -
 KF Tirana vs KF Teuta Durrës (26/04/2023) Semi-final Kupa e Shqiperise PES 202113 junho 2024
KF Tirana vs KF Teuta Durrës (26/04/2023) Semi-final Kupa e Shqiperise PES 202113 junho 2024 -
Mike ABC on X: drawing of the final battle of scp 1730 or What13 junho 2024
-
 Isekai Nonbiri Nouka - Episódio 3 - Animes Online13 junho 2024
Isekai Nonbiri Nouka - Episódio 3 - Animes Online13 junho 2024 -
 GTA 3 PS2 Gameplay13 junho 2024
GTA 3 PS2 Gameplay13 junho 2024
